Differential Expression Analysis
Differential Expression Analysis (DEA) performs statistical testing to determine if a given difference in read counts, between conditions, is significant or just due to random variations. You may run DEA at the gene, protein, and transcript levels, see image below. Protein and gene expression levels are calculated using the sum of their corresponding normalized transcript expression levels. You may also choose which R package to use, NOISeq or edgeR.
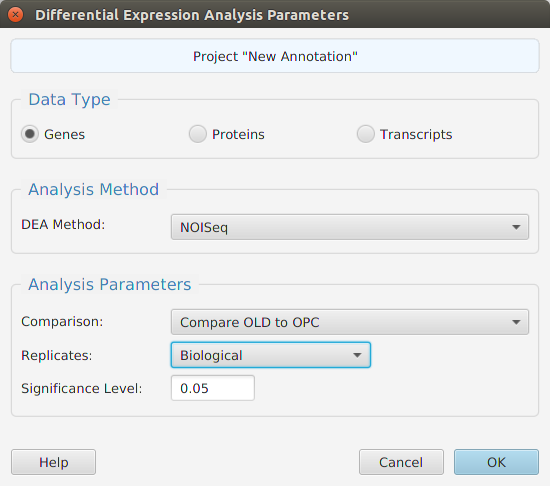
When using the application, all DEA parameters are described in the Help page which can be accessed via the Help button located on the bottom left of the dialog window.
NOISeq
NOISeq provides “differential expression between two experimental conditions with no parametric assumptions”.You may view the documentation and installation instructions at:
edgeR
EdgeR provides “differential expression analysis of RNA-seq expression profiles with biological replication. Implements a range of statistical methodology based on the negative binomial distributions, including empirical Bayes estimation, exact tests, generalized linear models and quasi-likelihood tests”.You may view the documentation and installation instructions at:
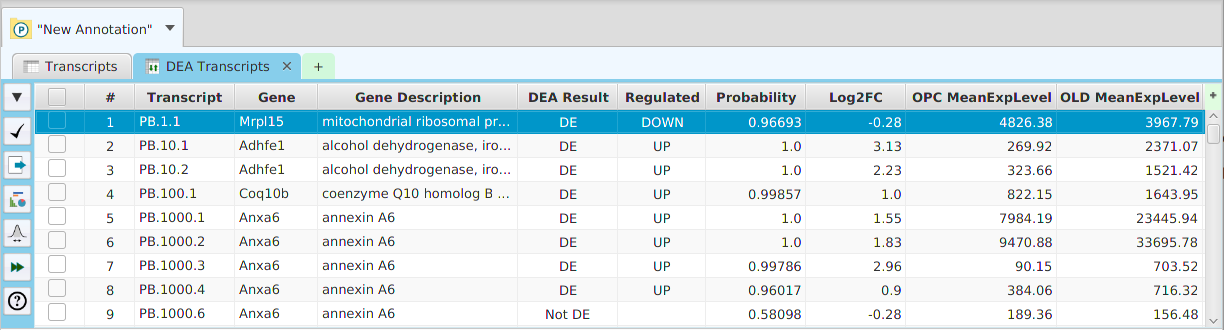
DEA Results
The DEA results are displayed in a table on the DEA Results subtab. The subtab is contained in the project data tab located in the top tab panel, see image below. It includes basic informational fields about gene, transcripts, or proteins depending on the DEA data type selected. It also includes the DEA test results, DE/NotDE and Up/Down regulation if DE, Probability or P-Value, depending on the R package used, and the Log2 of the fold change (Log2FC). The mean of the normalized expression levels for each condition are shown for each row.
Be aware that you do not need to rerun the analysis to change the significance level value: a menu button is provided in the subtab menu bar, left part of the image, to change the significance level value and recalculate the DE/NotDE results. When running the application, a description of all fields in the result table can be viewed using the subtab Help button.