Differential Isoform Usage
Differential splicing analysis can be performed for transcripts or proteins, see image below. Just like in the normal DIU using transcripts, using proteins allows checking for differential splicing at the protein level. Protein levels for DIU are calculated using the sum of their corresponding normalized transcript expression levels within the same gene. You may choose which R package to use, DEXSeqor edgeR, for DIU Analysis.
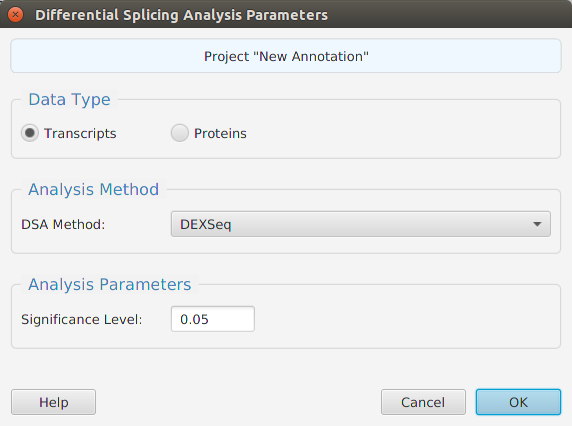
When using the application, all DIU parameters are described in the Help page which can be accessed via the Help button located on the bottom left of the dialog window. Also, be aware that you do not need to rerun the analysis to change the significance level value: a menu button is provided in the subtab menu bar, see Subtabs Menu Bar section, to change the significance level value and recalculate the DS/NotDS results.
DEXSeq
DEXSeq provides “…”.You may view the documentation and installation instructions at:
edgeR
EdgeR provides “…”.You may view the documentation and installation instructions at:
DIU Results
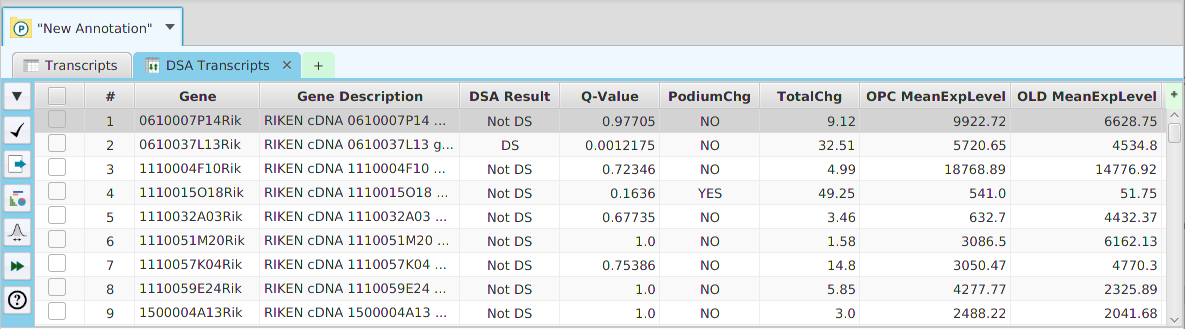
The DIU results are displayed in a table on the DIU Results subtab. The subtab is contained in the project data tab located in the top tab panel, see image below. It includes basic informational fields such as gene and gene description. It also includes the results, DS/NotDS, Q-Value or P-Value, depending on the R package used, Total Change, and Podium Change. Podium change is used to indicate if the most expressed transcript or protein, depending on the data type selected, changed between conditions. The mean normalized expression levels for each condition are shown for each row.
Be aware that you do not need to rerun the analysis to change the significance level value: a menu button is provided in the subtab menu bar, left part of the image, to change the significance level value and recalculate the DIU/NotDIU results. When running the application, a description of all fields in the result table can be viewed using the subtab Help button.